Paul Hegedus
Research Overview:

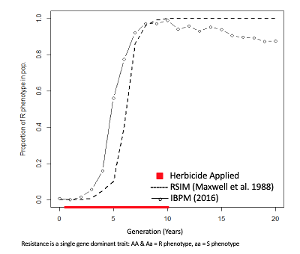
Herbicide resistant weeds are an increasing threat to agricultural systems around the world. The reliance of high input agriculture on pesticides has led to the evolution of herbicide resistance in over 250 species, some of which are resistant to multiple herbicide’s site of action. Herbicide resistance modeling has been around for decades, most often in the form of population-based models. These models treat all the individuals in a population as a unit while individual-based models act independently on each individual in the population. The strength of individual-based models is their ability to address and express the realism of biological systems. This research involves developing a generic individual-based herbicide resistant simulation model to determine the variation in resistant phenotype dynamics for a single dominant allele mechanism. From this work, the rate of evolution of resistance in an individual-based model has been found to be greater compared to a population-based model. Additionally, the individual-based model indicates the potential for resistance to decline in generations after a peak proportion of resistance is achieved in the population. This is likely due to mating between heterozygotes and/or homozygous recessive individuals that reintroduce the susceptible phenotype to the population. Further objectives for this research include assessing the range of resistance evolution outcomes under different assumptions about 1) the initial frequency of resistance in the population, 2) the initial number of patches of individuals, 3) the management strategy applied to patches, 4) the mating behavior of individuals, and 5) fitness costs associated with resistant alleles.