Accelerating the development of barley varieties for improved forage quality and yield
Traci Hoogland: MS Candidate
Problem Statement:
“Depending on location, producers need a 2 to 4 month supply of hay to get through the winters in the northern Great Plains. Aside from long periods of snow cover, high-quality forages are required to offset poor-quality roughages available on range. Winter feed is the largest cost on ranching operations, and slight improvements in forage production can significantly reduce costs.” Dr. Dennis Cash, MSU Forage Extension Specialist for 14 years (2006)
Barley, a highly drought and saline tolerant crop, is uniquely suited to hay production in Montana and an important source of winter feed for ranchers across the state. Developing and releasing barley hay varieties with higher quality and biomass yields will benefit our growers and ranchers, but breeding for these traits with only conventional tools is expensive and may require as many as 8-10 years to develop a single new variety.
Purpose:
The purpose of this project is to accelerate the development of Montana-adapted barley varieties with improved hay quality and yield.
Objectives:
1) Utilize near-infrared reflectance (NIR) technology to make screening for forage quality faster and cheaper – allowing more samples, and thus more barley lines across more environments, to be tested
2) Identify germplasm with superior digestibility and biomass yield for incorporation into the MSU barley breeding program
3) Improve our understanding of the genetic variation underlying forage digestibility
o Discover novel QTLs (Quantitative Trait Loci) related to digestibility and biomass yield
o Utilize bioinformatics to interrogate QTL regions for candidate genes for additional study
4) Develop breeder-friendly genetic markers associated with forage digestibility and yield for Marker-Assisted Selection (MAS)
Hypothesis:
Germplasm resources such as the USDA Barley Core Collection (BCC), maintained by the National Plant Germplasm System, are an incredible, largely untapped repository of genetic diversity.
A genome-wide association study utilizing a subset of the BCC will allow us to uncover novel QTLs associated with the traits of interest, while simultaneously identifying lines with superior forage traits which can then be crossed into Montana adapted lines within the MSU barley breeding program.

Project Description:
Background:
Forage quality has three major components:
· Intake: relates to an animal’s desire and ability to consume a given forage
· Digestibility: relates to an animal’s ability to break down and utilize nutrition from a forage it has consumed
· Nutrition profile: relates to what an animal is able to get from a forage once it is digested
Here we focus on intake and digestibility
Intake and Digestibility are estimated via laboratory analysis:
Forage Intake is measured via a Neutral Detergent Fiber (NDF) test:
· The portion of a forage sample remaining after digestion in a detergent with a neutral pH. This value is negatively correlated with forage intake.
Forage Digestibility is measured via an Acid Detergent Fiber (ADF) test:
· The portion of a forage sample remaining after digestion in a detergent with an acidic pH predicts digestibility.
· As digestibility increases, livestock average daily gains increase.
In cattle, a 1% increase in digestibility has been shown to lead to a 3% increase in average daily gains.
(Casler et al. 1999, Suber et al. 2003 unpublished data, Mohammed et al. 1967)
Research Techniques:
· Genome-Wide Association Mapping (GWAS)
o 260 genotyped lines were selected from the BCC based on contributed genetic diversity
o Lines were grown in an augmented block design in Bozeman, MT under both dryland and irrigated conditions
o Lines were phenotyped for forage quality, biomass yield, and other key agronomic traits
o By comparing variation in forage quality traits to variation in genome-wide genetic markers, a mathematical model can be used to find associations between forage traits and the genetic regions impacting these traits
Results
 |
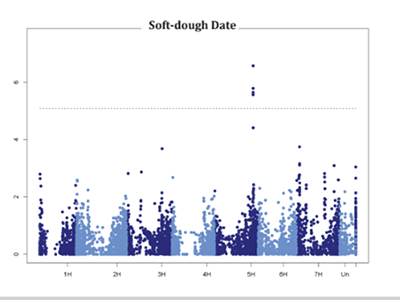 |
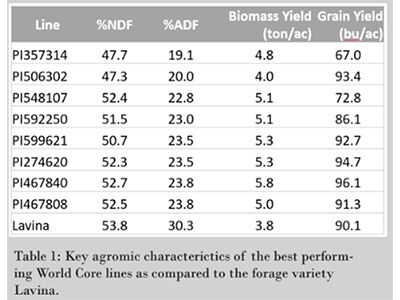
| Many lines were identified which had higher estimated forage yield and quality than Lavina, one of the most commonly grown hay barley varieties in Montana. The top performing lines identified in the 2016 field season (Table 1) were immediately added to the forage barley breeding program | Lines were monitored daily through-out the growing season and forage sampling was conducted on the day a line reached the soft-dough stage of maturity. One of the QTLs identified in a preliminary analysis of the 2016 data was associated with the soft-dough trait. A Manhattan plot of this marker-trait association is displayed here. |
Utilization of NIR technology
o NDF and ADF values were collected on more than 200 barley forage samples, these samples were then used to develop a customized NIR calibration curve
o With this NIR technology, we have been able to collected forage quality data on more than 1200 forage samples with a fraction of the time and cost of other analytical methods


