Genetic Dissection of Malt Quality
Joeseph Jensen: PhD Candidate
Problem:
- For a malt house time is money especially since the malting process is very resource intensive, requiring water, energy, and a weeks’ worth of processing to produce a single batch of malted grain. If the malting process could be shortened it could save a malt house money. One way we have found that could reduce malt times is by increasing the rate at which water enters the grain and hydrates the endosperm, also know as the speed of hydration. The question is what affects will this have on malt quality of the grain.
- From the grower’s perspective improving our understanding of speed of hydration will also be beneficial. Historically faster hydrating lines lacked dormancy. If lines have dormancy, they are more resilient to preharvest sprouting during an untimely rain event. We theorize that by understanding the genetic controls of dormancy and speed of hydration we can breed lines that are fast to hydrate and have dormancy so that they will meet both growers and maltsters needs.

Purpose/Objectives:
- By understanding the genetic controls of malt quality, speed of hydration and dormancy we can breed lines that have some dormancy, fast hydrating and have good malt quality. This stacking of traits will also provide growers with a toll to prevent pre harvest sprouting.
Hypothesis:
- We hypothesize that dormancy and speed of hydration are independent traits.

Project description:
- This work is being done using two mapping populations. The first is a genetically diverse population of 169 modern and historic malting lines selected form GRIN. With this population we are using association mapping to identify QTLs for our desired traits. With this population we will be able to find genes that may be fixed in a modern malting barley population. The second population was derived from a cross between two modern malting barley lines MT124128 and MT124148. With this population we are using biparental mapping to identify QTLs. The advantage with using this type of genetic mapping is we will be able to find small affect QTLs that you would not normally be able to identify using association mapping. So, by using both population to find speed of hydration QTLs we should be able to find a wide variety that we can use for breeding.
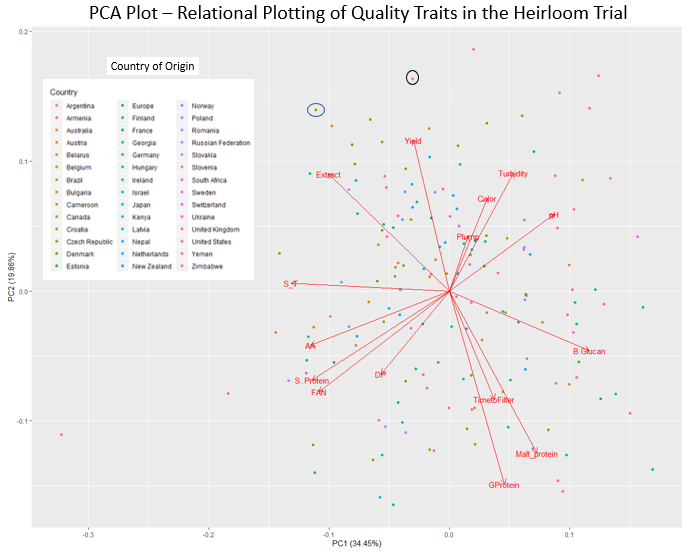 PCA plot grouping varieties based on quality performance. Each arrow identifies an
important malt quality trait. The arrows point towards the varieties with highest
value for the given trait. Note arrows for extract and yield are pointing in the
same general direction, indicating highest yielding lines are also highest extract.
Arrows for yield and extract are pointing in opposite directions of grain protein
and beta glucan, meaning as yield and extract increase grain protein and beta glucan
decrease. However, the scatter of the varieties across the plot indicate that while
these trends exist, some lines fit the trend better than others. The circled varieties
would have high yield and extract but low protein and beta glucan. This graph allows
us to consider all the traits simultaneously. For example, a brewer might want a
line with less FAN. In that case we would select the black circled line.
PCA plot grouping varieties based on quality performance. Each arrow identifies an
important malt quality trait. The arrows point towards the varieties with highest
value for the given trait. Note arrows for extract and yield are pointing in the
same general direction, indicating highest yielding lines are also highest extract.
Arrows for yield and extract are pointing in opposite directions of grain protein
and beta glucan, meaning as yield and extract increase grain protein and beta glucan
decrease. However, the scatter of the varieties across the plot indicate that while
these trends exist, some lines fit the trend better than others. The circled varieties
would have high yield and extract but low protein and beta glucan. This graph allows
us to consider all the traits simultaneously. For example, a brewer might want a
line with less FAN. In that case we would select the black circled line.


